Overview
smFRET reports nanometer-scale donor/acceptor distance changes in single molecules, enabling direct observation of conformational switching kinetics in living-cell conditions. In this repository, Hsp90 dynamics are represented as a three-state chain: Open (O), Intermediate (I), and Closed (C).
State topology
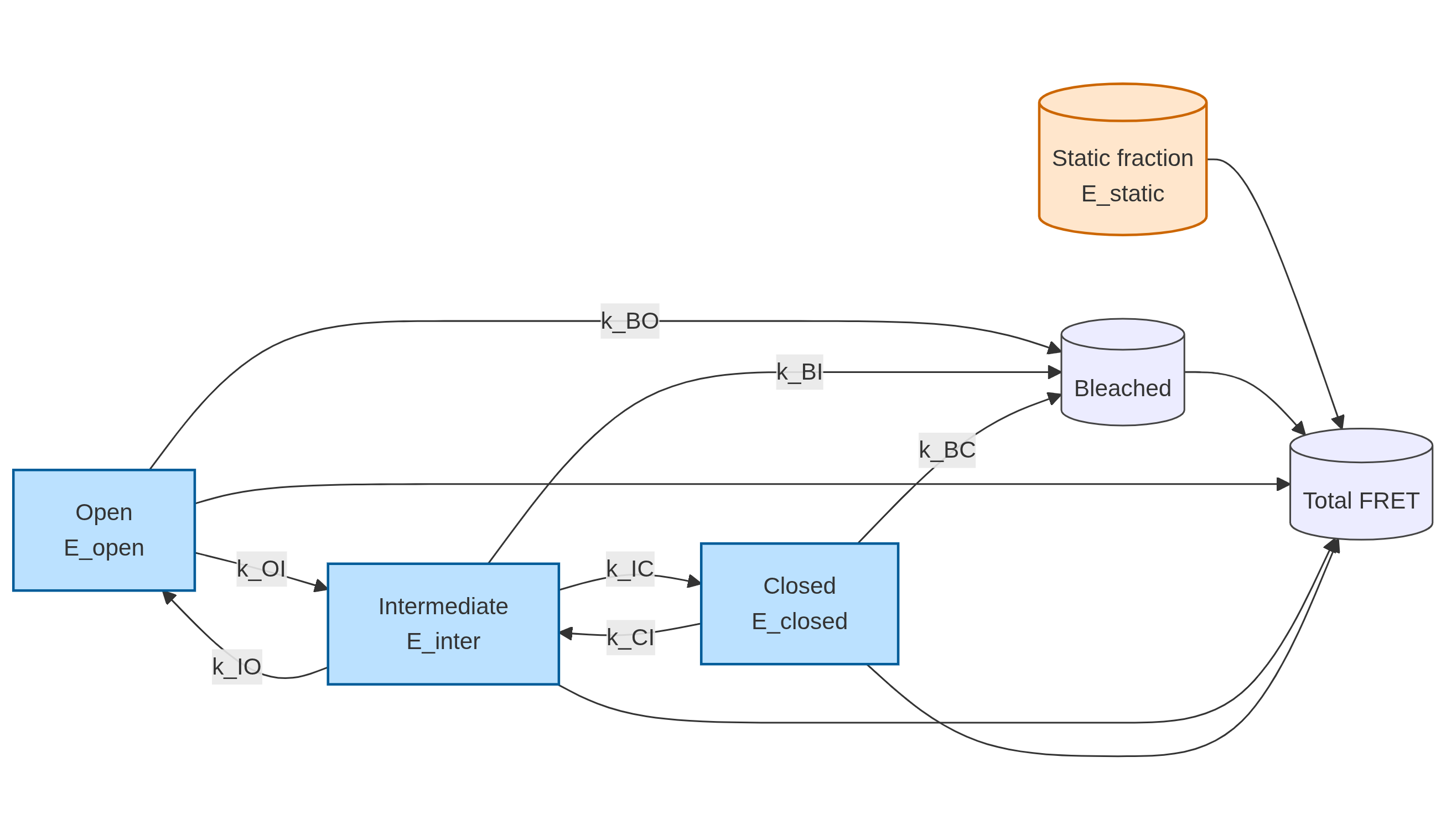
Dynamic + static mixture
The ensemble signal is modeled as a mixture of:
- a dynamic population with fraction
f_dyn, evolving through the O↔I↔C network plus bleaching, and - a static population with fixed FRET level
E_static.
This separation allows the fit to capture both kinetic transitions and non-switching subpopulations.
Processing flow
flowchart LR
A[.h5/.tracks] --> B[get_traces.py]
B --> C[fret_matrix.csv]
C --> D[pipeline.py]
D --> E[results/]
Note
The full analysis depends on the Hügel 2025 dataset (Zenodo DOI: 10.5281/zenodo.17559063). Place downloaded files in data/Hugel_2025/ before running the pipeline.